http://english.cas.cn/newsroom/research_news/life/202508/t20250801_1048868.shtml
https://biotechnologyforbiofuels.biomedcentral.com/articles/10.1186/s13068-025-02677-8
A team at CAS Qingdao Institute of Bioenergy and Bioprocess Technology (QIBEBT) has developed a lipid-rich mutant strain of Saccharomyces cerevisiae for microbial production of palmitoleic acid— a rare omega-7 fatty acid with proven anti-inflammatory and metabolic benefits.
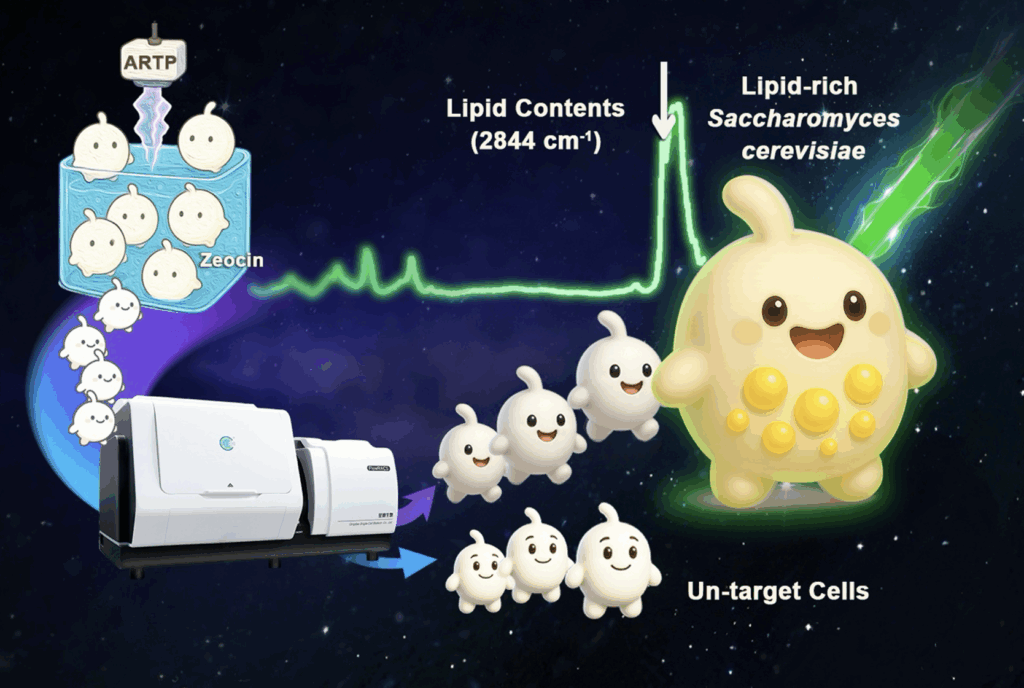
The team used a combined mutagenesis approach—employing zeocin, an antibiotic-based mutagen, and Atmospheric and Room Temperature Plasma (ARTP)—to create a diverse library of yeast mutants. They then deployed FlowRACS, a Raman flow cytometry system, to select live yeast cells with elevated lipid levels by analyzing their intrinsic single-cell Raman spectra, eliminating the need for chemical stains or genetic reporters.
This method identified the mutant strain MU2R48, which achieved a lipid content of 40.26%—a 30.85% increase over its parental strain SC018—while maintaining similar biomass production.
Photo: Raman flow cytometry efficiently identifies lipid-rich Saccharomyces cerevisiae mutants from a Zeocin–ARTP-induced library. (Image by QIBEBT)