http://english.qibebt.cas.cn/ne/rp/202504/t20250407_909473.html
https://pubs.acs.org/doi/10.1021/jacs.4c18730
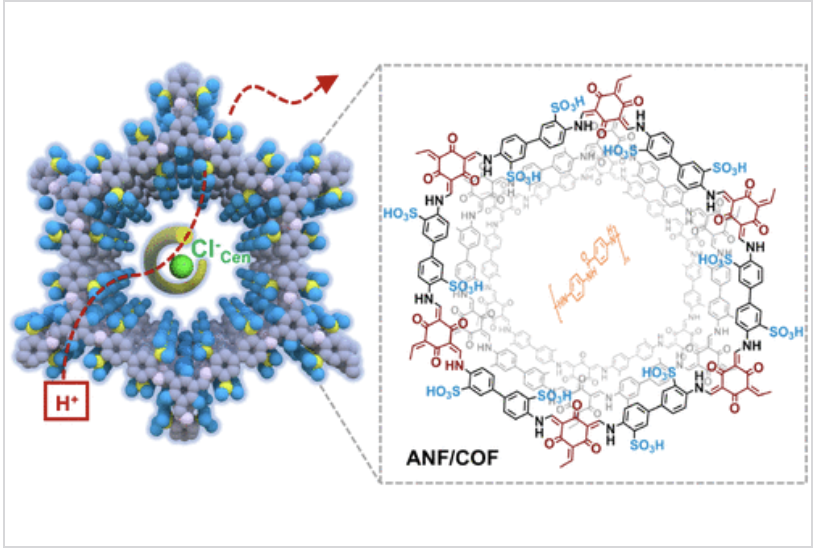
A research team from the CAS Qingdao Institute of Bioenergy and Bioprocess Technology (QIBEBT) has introduced a novel membrane design that mimics biological protein channels to enhance proton transport for efficient energy harvesting. Inspired by the ClC-ec1 antiporter found in Escherichia coli, which facilitates the movement of chloride (Cl⁻) and protons, the researchers developed a hybrid membrane composed of covalent organic frameworks (COFs) integrated with aramid nanofibers (ANFs). This ANF/COF composite forms a robust hydrogen-bonding network and features amide groups that selectively bind to Cl⁻ ions, significantly lowering the energy barrier for proton conduction.
In acidic environments, adding just 0.1% Cl⁻ ions (relative to protons) increased the membrane’s proton permeation rate threefold, reaching 9.8 mol m⁻² h⁻¹ for the efficient migration of H⁺ ions. Under simulated acidic wastewater conditions, the ANF/COF membrane achieved an output power density of 434.8 W m⁻²—one of the highest reported to date for osmotic energy generation. It also showed structural stability over 9,000 minutes (~150 hours) of operation in highly acidic media.